Pair-distance distribution analysis – BIFT in RAW
RAW has a built in method for determining the P(r) function using a Bayesian IFT method (BIFT). This has the advantage of only have one possible solution. More information on this method can be found in the RAW paper and manual and references therein.
This tutorial covers how to use RAW for doing an IFT. This is not a tutorial on basic principles and best practices for doing an IFT or analysis of the resulting P(r) function. For that, please see the SAXS tutorial.
If you use RAW to run BIFT, in addition to citing the RAW paper, please cite the BIFT paper: Hansen, S. Journal of Applied Crystallography (2000) 33, 1415-1421. DOI: 10.1107/S0021889800012930
A video version of this tutorial is available:
The written version of the tutorial follows.
Right click on the glucose isomerase profile in the Profiles list you loaded previously. Select “IFT (BIFT)” from the resulting menu.
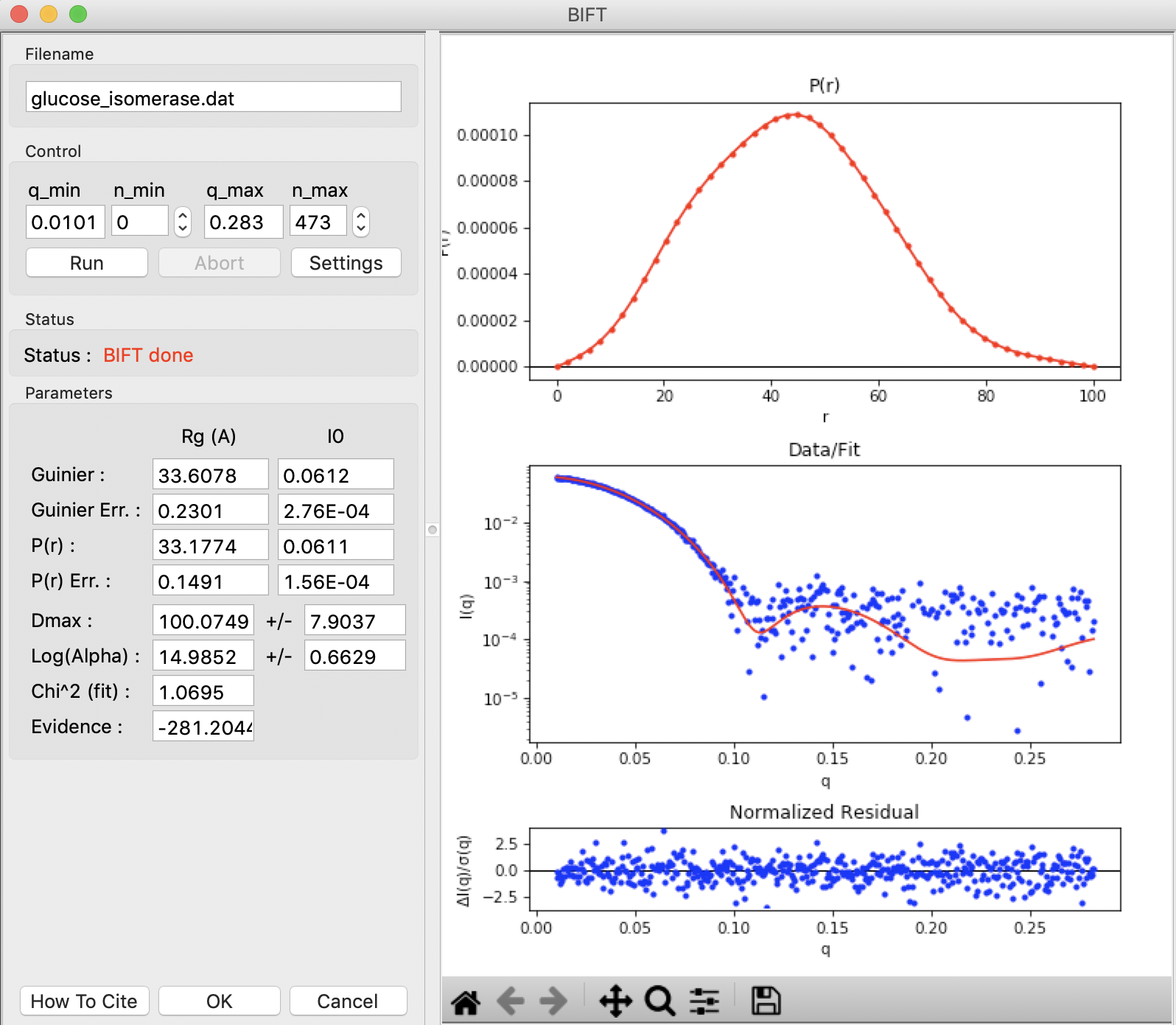
The BIFT panel has plots on the right. These show the P(r) function (top panel), the data (middle panel, blue points) and the fit line (middle panel, red line), and the fit residual (bottom panel).
On the left of the BIFT panel are the controls and the resulting parameters. Note that in this case you do not control the Dmax value, the BIFT method finds that for you automatically. Because BIFT can take some time to run, if you change the q range for the data you have to click the ‘Run’ button to run BIFT again.
Note that for this dataset, BIFT has found a Dmax value around 100, in good agreement with what we found from GNOM.
Click OK to exit the BIFT window. This saves the results into the IFTs panel.
Click on the IFTs Control and Plot tabs. This will display the BIFT output you just generated. Save the glucose_isomerase.ift item in the reconstruction_data folder.
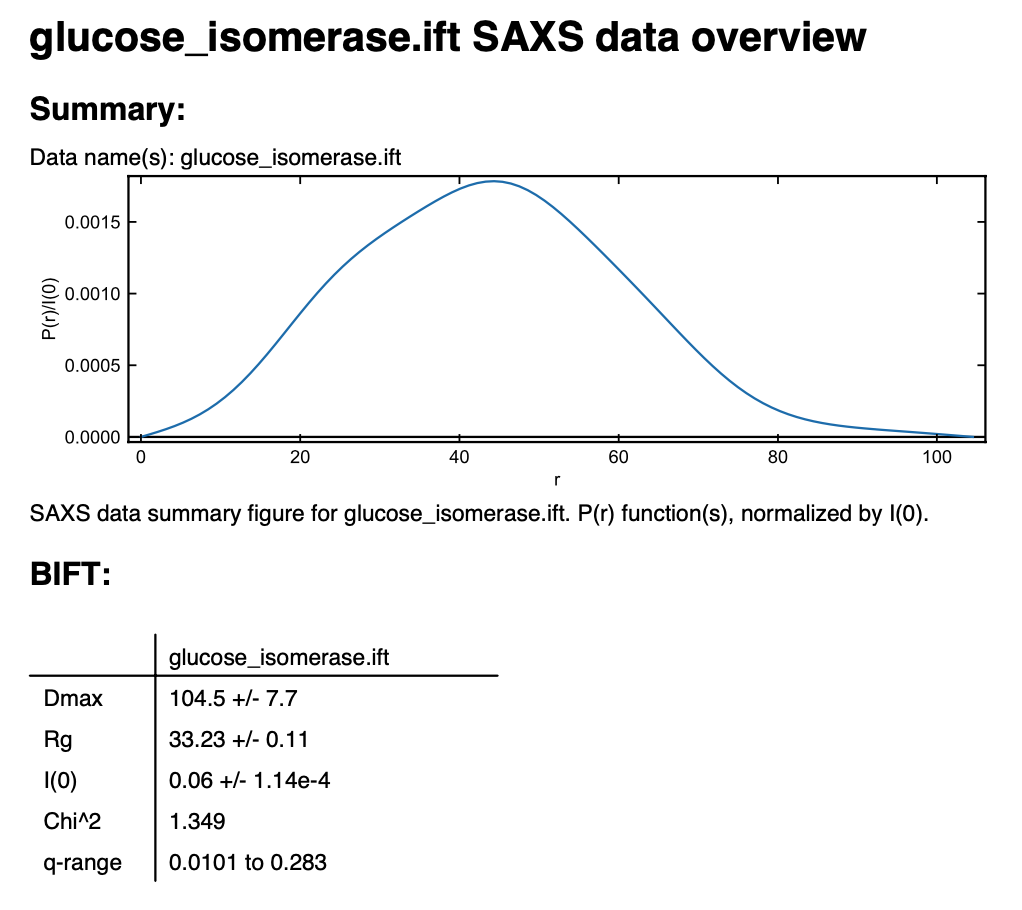
You can save the IFT in the pdf report that RAW can make. Right click on the glucose_isomerase.ift item in the IFT control panel and select “Save report”. In the window that opens click “Save Report” and save the pdf report. If you open the report you will see a plot of the P(r) function and a summary of the run parameters and numerical results saved as a table.
Note: As of now, BIFT output from RAW is not compatible with DAMMIF or other ATSAS programs. However, it is compatible with electron density determination via DENSS.